An article from University of Oslo

Bacterial DNA from the Black Death found in teeth
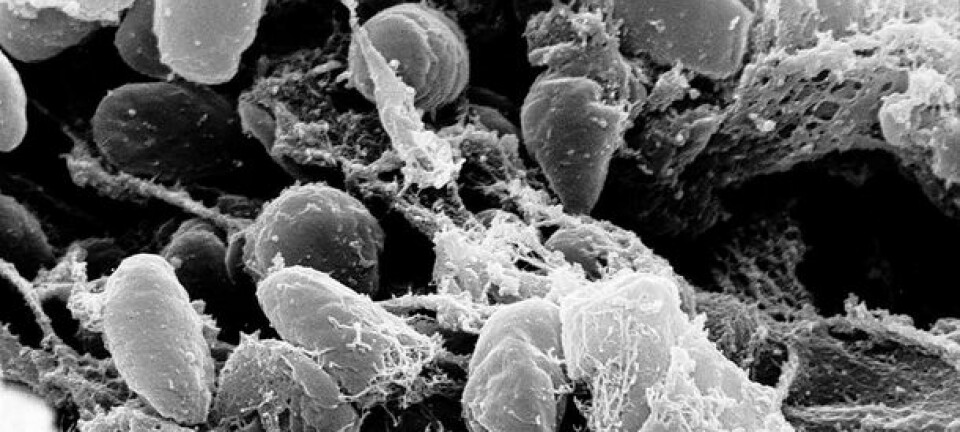
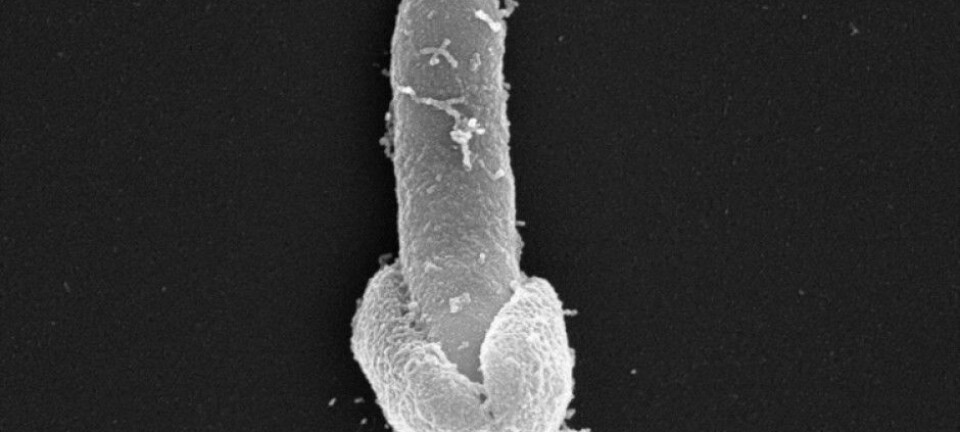
Remnants of the genetic makeup of plague bacteria have been found in thousands of victims of the Black Death and the major plague epidemics at the end of the Iron Age. The DNA analyses may predict the next plague outbreak.
Denne artikkelen er over ti år gammel og kan inneholde utdatert informasjon.
Biologists at the University of Oslo are now making a giant effort to identify the relationship between climate change, rat infestations, and the many major plague epidemics throughout history. The knowledge may be used to predict the next plague outbreak.
Plague did not only occur in the Middle Ages. There are 2,000 cases of plague annually; most of them in Madagascar and Congo. Outside Africa, plague occurs in the deserts of North America and in large areas in Central Asia, in a wide belt from Georgia via Kazakhstan to China.
The world has suffered three major and many small plague epidemics in the Common Era. The first major pandemic was the Justinian Plague at the end of the Iron Age, which lasted for 200 years. The next apocalypse was the Black Death, a pandemic that kept coming in new waves, and lasted for 400 years.
The third epidemic, which began at the end of the 19th century in China and reached the rest of the world via Hong Kong, is still ongoing. This summer a man died from plague in the city of Yumen in northern China. 30,000 inhabitants were confined to isolation when parts of the city were locked down. There was a plague outbreak in November in Madagascar, both in the provinces and in the capital of Antananarivo.
The evolution of plague bacteria

The big question is where the plague bacterium comes from, whether the three plagues stem from the same bacterium, and how the plague bacterium survived between the outbreaks. Some people believe that the plagues at the end of the Iron Age and during the Middle Ages were two different diseases, but several studies show that they were the same disease.
"The transitions between the three plague outbreaks are not completely clear. There was also an overlap between the second and the third outbreaks. Plague can completely vanish for decades, and then return," says Professor Kjetill S. Jakobsen of the Centre for Ecological and Evolutionary Synthesis (CEES), University of Oslo.
The question is: in which circumstances does plague become pandemic and what determines the level of severity of the epidemic?
"It is tempting to speculate whether the ecology of the locations in which the pandemics started played a role in their level of destruction. We can understand the serious pandemics in the past, present and future by looking at the link between climate conditions and plague dynamics," says Professor Nils Christian Stenseth, Chair of CEES.
Examining old skeletons
The understanding of the three plague pandemics has improved vastly during the past few years because researchers can now use modern molecular methods. This allows the analysis of DNA remnants of plague bacteria from plague victims.
Italian researcher Barbara Bramati of CEES is studying the evolutionary development of the plague bacteria, and is comparing the genetic code of the plague bacteria during the first two pandemics.
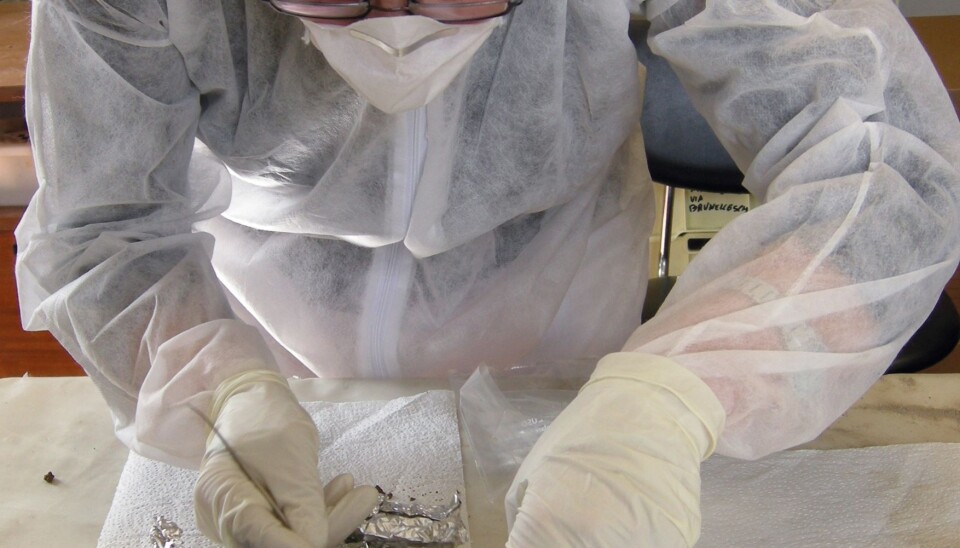
She is finding her answers in the teeth of old skeletons. The lymph nodes of the plague victims swelled up and transformed into black boils, where the bacteria would multiply. In many of the victims, the plague bacteria entered the bloodstream, and thus the dental pulp. Together with archaeologists, Bramanti is collecting DNA samples from the teeth of 2,700 plague victims from Europe and Asia. She will also be checking the teeth of Norwegian victims.
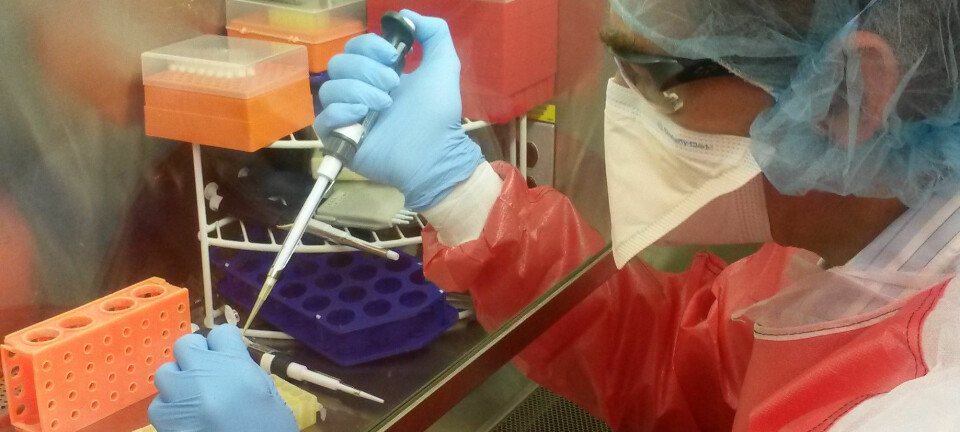
She drills a few millimetres into the teeth and extracts the entire DNA content of the dental pulp.
"The DNA remnants are often fragmented and destroyed. This makes reconstructing the genetic material an arduous task."
The Department of Biosciences at UiO is now opening a completely new DNA laboratory that will make it easier to analyze the DNA remnants in detritus. The laboratory will be the largest of its kind in Europe.
"Here it will be possible to analyze even small quantities of ancient DNA."
As long as the quality of the genetic material is good enough, she will compare it with the reference genome of the plague bacterium.
"Our point is to look at both the evolution and the ecology of the plague bacterium," says Barbara Bramanti.
Examining rodents
It is not clear how the Black Death spread. Black rats and brown rats spread contagion during the third plague outbreak. Even though there still is no proof that gerbils were carriers, gerbils appear to handle the contagion surprisingly well.
"There are great individual differences, but many individuals can handle an absurd amount of plague bacteria," says PhD research fellow Pernille Nilsson at CEES. She has studied how much gerbils can tolerate.
The test was carried out at a plague laboratory in China, not far from Yumen, which had a case of plague this summer. The researchers injected so many plague bacteria into the rodents that nearly half of them died. This made it possible for them to compare the genome of the individuals that survived with the genome of the dead.
In other words, Nilsson is looking for the genetic explanation as to why the immune systems of gerbils are so strong against the plague bacterium.
"Sometimes a single bacterium kills a mouse. Common rats can tolerate injection of 10,000 bacteria. Gerbils can tolerate 100 billion bacteria. That is ten million times as many bacteria," says Pernille Nilsson.
To achieve the best possible DNA analysis of each gerbil, she must also compare it with the fully-sequenced genome of the gerbil species. Enormous work is involved in fully sequencing the genome. It took ten years to sequence the human genome. When Jakobsen's research group identified the genome of cod, the DNA sequencing only took a few months, with a couple of years of computationally-intensive DNA calculations behind them.
Thanks to the much faster laboratory machines and computer programs of the present, Nilsson hopes to fully sequence the gerbil's genome much faster. For now she will have to deal with the partly-sequenced genome.
China has already extracted and freeze-dried DNA from the livers and spleens of three different gerbils. Researchers at UiO then checked the quality of the DNA before they decided which individual to collect the reference genome from.
The genome of a gerbil consists of 2.4 billion base pairs. When the genome is sequenced, only one hundred base pairs can be read at a time. To determine the location of the small nucleotides in the genome, they must be processed a number of times and huge computational analyses must be run to sequence them correctly. And every time they compare the genome from a single gerbil with the fully-sequenced gerbil genome, they need one week of access to the university's supercomputer.
"We must find the regions in the genome where there is natural variation among individuals and the regions of the genome that show specific differences between the gerbils that survive and the ones that die. We have not identified these regions in the genome yet," says Nilsson.
The researchers believe that protection against plague is related to a congenital immune system and that this is hereditary.
"We do not know quite what it is yet. We are on a 'genomic fishing expedition', and don't quite know what we will find. We still have some ideas as to which parts of the immune system that may be involved."
A few years ago, CEES discovered that cod have a completely different immune system than all other animals.
"We may also make some surprising discoveries about the immune system of gerbils," says Nilsson.
Climate link
Dutch researcher Boris Schmid of CEES is studying the relationship between climate variations, the rodent population and plague outbreaks. He combines historical time series with plague data from humans and the relationship between climate change and plague-stricken rodents.
China has also run DNA sequences of the plague bacterium for 20–30 years. This makes it possible to see how the plague bacterium has developed in evolutionary terms during this period.
"This data will give us an idea as to how plague and the climate are linked, and is important for us to be able to predict the next plague outbreak," says Boris Schmid.
--------------------------------------------
Read the Norwegian version of this article at forskning.no
External links
- Kjetill Sigurd Jakobsen's profile
- Nils Christian Stenseth's profile
- Barbara Bramanti's profile
- Pernille Nilsson's profile